Authors: Markus Wilken and Sikelela Buthelezi
Forestry and Agricultural Biotechnology Institute FABI
Department of Biochemistry, Genetics and Microbiology, University of Pretoria.
Introduction
Sclerotinia sclerotiorum is a fungal pathogen of many crop species that needs no introduction to anyone working in the Agricultural industry. In South Africa, S. sclerotiorum causes disease on many economically important crop species including canola, soybean and sunflower. The persistent occurrence of this fungus in the field is attributed to its ability to form hardened, long-term survival structures called sclerotia, a trait it shares with other plant pathogens (e.g., Claviceps purpurea causing ergot on cereal crops) and even some mushroom species (e.g., Psilocybe mexicana, the psychedelic mushroom). Sclerotia of S. sclerotiorum form the basis for its infection cycle, with severe outbreaks leading to significant yield losses.
Its notoriety as an economically important pathogen has caused S. sclerotiorum to be a well-researched fungus. Although much is known about its infection cycle, the ability to cause disease in a range of plants is still not well understood, which is a rather usual property among plant pathogens. This has led to many studies focussing on the way in which S. sclerotiorum causes disease and identifying a wide array of biological molecules that affect its pathogenicity, as well as ways to control it in the field. For example, viruses 1 and antagonistic fungal species 2 that adversely affect S. sclerotiorum have been identified. These studies have boosted our understanding of the pathogen significantly, but as is commonly the case, the elusive “silver bullet” has not been found.
Genomes & Genetic Instructions
Research is intricately linked to technology, and following the development of so-called next generation sequencing platforms3 we now have access to the full genome sequences of many living things. A genome sequence is a copy of all the genetic instructions present in a cell, and essentially represents the blueprint on what makes a species what it is. The availability of full genome sequences has thus impacted all aspects of biology, including modern plant pathology where it has provided a plethora of new tools to use for enhancing plant health. For S. sclerotiorum (strain 1980), a high-quality draft of its genome sequence was produced in 20054 and, by using newer sequencing technologies, was completed in 20175. The S. sclerotiorum genome has 38,7 million nucleotides, which is about 1.5% the size of the human genome. Despite its small size, the S. sclerotiorum genome contains roughly 50% as many genes found in humans. However, the functions of most of these genes are not yet known, highlighting the complexity that underlies the functioning of something seen as a “simple” fungus.
The tools provided by a genome sequence is useful for many studies related to fungal pathogens. Genomes offer the chance to develop population-level markers specific for a target species6. This has significant benefits for epidemiological studies in that it allows estimation of genetic diversity and modelling future infection risks. A genome sequence also provides a catalogue of the history of a species that can be picked apart for clues to the origin and biological functioning of the fungus7. Although sounding very simple, these are complex tasks that require huge contributions from many scientific fields.
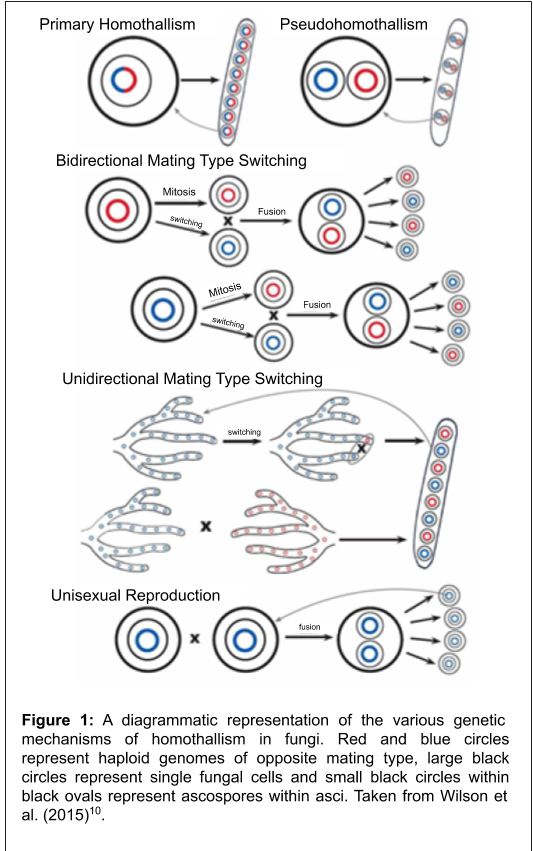
One study area that has benefitted from the availability of genome sequences is research into the reproductive strategies of fungal pathogens8. The sexual process in fungi is controlled by genes present at a specific location in the genome. Knowledge of the structure of this region provides insight into how these fungi reproduce, which in turn has important implications on the population dynamics of a species9. A species that employs an exclusively outcrossing or heterothallic reproductive mode (Figure 1) will need to find a compatible mating partner and subsequently produce genetically diverse progeny that can increase the adaptability of a species. In these fungi, if a compatible mating partner is not available, sexual reproduction will not occur, which could have broad impacts on the population genetics of the fungus. By contrast, in homothallic species (Figure 1), mating can occur in the absence of a partner. Here, an isolate can initiate sexual reproduction in solo and produce thousands of sexual progeny by itself, resulting in a population of clones – individuals that are genetically identical. However, they retain the ability to outcross, and can thus benefit from outcrossing and the generation of variation similar to heterothallic species.
Sclerotinia sclerotiorum is homothallic and can therefore reproduce in the absence of a mating partner. During this process, the fungus produces the characteristic mushroom-shaped apothecia, which releases sexual spores that can establish an infection on the plant. As expected, many field populations of the fungus show high levels of clonality due to homothallism. However, some populations studies show moderate levels of genetic diversity and the genetic signatures of outcrossing, putting the reproductive strategy of this fungus under the microscope11. From a molecular viewpoint, the sexual process is even more bizarre as it incorporates a change at the DNA level that structurally and functionally influences the sexual process12. This leads to questions on the link between the DNA rearrangement process and the occurrence of unexpected levels of genetic diversity in the field.
Our Research
Our project aims to establish a South African-specific genomic resource for studies on S. sclerotiorum. The availability of a high-quality draft genome from a South African isolate of the species will prove invaluable for expanding our current understanding of the pathogen in a local context. It would allow us to design genetic markers catered specifically to the South African population and target suitable regions for developing diagnostic tools, while at the same time providing insight into the evolutionary history of the pathogen in the country. With the genome information, we will also be able to study the reproductive strategy used by South African populations of the pathogen. Insights gained from these studies will not only add to the growing debate around the population genetics structure of the fungus, but also contribute meaningfully in efforts aimed at combatting this important pathogen in the field.
References
- Jiang, D., et al., Viruses of the plant pathogenic fungus Sclerotinia sclerotiorum. Advances in Virus Research, 2013. 86: p. 215-48.
- Smolińska, U. and B. Kowalska, Biological control of the soil-borne fungal pathogen Sclerotinia sclerotiorum –– a review. Journal of Plant Pathology, 2018. 100(1): p. 1-12.
- Schuster, S.C., Next-generation sequencing transforms today's biology. Nature Methods, 2008. 5(1): p. 16-18.
- Amselem, J., et al., Genomic analysis of the necrotrophic fungal pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS Genetics, 2011. 7(8): p. e1002230.
- Derbyshire, M., et al., The complete genome sequence of the phytopathogenic fungus Sclerotinia sclerotiorum reveals insights into the genome architecture of broad host range pathogens. Genome Biology and Evolution, 2017. 9(3): p. 593-618.
- Simpson, M.C., et al., Analysis of microsatellite markers in the genome of the plant pathogen Ceratocystis fimbriata. Fungal Biology, 2013. 117(7-8): p. 545-555.
- Sharma, K.K., Fungal genome sequencing: basic biology to biotechnology. Critical Reviews in Biotechnology, 2016. 36(4): p. 743-59.
- Wilken, P.M., et al., Which MAT gene? Pezizomycotina (Ascomycota) mating-type gene nomenclature reconsidered. Fungal Biology Reviews, 2017. 31(4): p. 199-211.
- Havenga, M., et al., Mating strategy and mating type distribution in six global populations of the Eucalyptus foliar pathogen Teratosphaeria destructans. Fungal Genetics and Biology, 2020. 137: p. 103350.
- Wilson, A.M., et al., Homothallism: an umbrella term for describing diverse sexual behaviours. IMA Fungus, 2015. 6: p. 207-214.
- Attanayake, R.N., L. Xu, and W. Chen, Sclerotinia sclerotiorum populations: clonal or recombining? Tropical Plant Pathology, 2019. 44(1): p. 23-31.
- Chitrampalam, P., et al., The Sclerotinia sclerotiorum mating type locus (MAT) contains a 3.6-kb region that is inverted in every meiotic generation. PLoS One, 2013. 8(2): p. e56895.